EGFR Mutation Detection Test
Improved sensitivity for single gene mutation detection
CE/IVD Marked Product
Research Use Product
Research Service
Early Detection/Screening
Diagnosis
Therapy Selection
Therapy Monitoring
Introducing QClamp® EGFR Test
The QClamp® EGFR Mutation Detection Test for Formalin-Fixed Paraffin-Embedded (FFPE) samples aids in the identification of patients eligible for cancer treatment and monitor response to therapy, which can lead to improved outcomes in cancer patients. It is an in vitro diagnostic real-time quantitative PCR assay for the detection of somatic mutations in and near codon 719 in Exon 18, Exon 19 deletions, codons 790, and insertions in Exon 20, and codons 858 and 861 in Exon 21 in the human EGFR gene, using purified DNA extracted from FFPE (for CE/IVD version) or plasma (for research use version).
With proper validation, the kit can be used for low-frequency mutation detection of important EGFR mutations such as resistance mutation T790M using plasma samples.
Ultra-sensitive: reliably detects 0.1% to 0.5% VAF mutant DNA out of wild-type DNA for targeted mutations
Low DNA input: Minimum 5ng input DNA per reaction for RUO (research use) and 10 ng DNA per reaction for CE/IVD
Fast result: Less than 4 hours of assay run time
Validated on the most common qPCR machines with minimized variability
EGFR Mutation and Lung Cancer
EGFR Introduction
The epidermal growth factor receptor (EGFR) is a membrane protein that possesses an intracellular receptor tyrosine kinase (TK) on the surface of epidermal cells. EGFR plays a central role in transmitting signals that promote cell growth, survival and proliferation by activating several downstream pathways including the Ras-Raf-MAPK pathway, the PI3-kinase-PDK1-Akt pathway, and JAK1/2-STAT pathways. EGFR overexpression and presence of oncogenic mutations that constitutively activate the TK domain of EGFR have been found in various solid tumors such as lung cancer and colorectal cancers. For instance, some of the KRAS mutations are associated with resistance to TK inhibitors, erlotinibor and gefitinib, in lung adenocarcinoma patients with EFGR mutations.
Target Therapy for EGFR
The most common activating mutations in the EGFR oncogene, exon 19 deletion and L858R, respond to certain tyrosine kinase inhibitor (TKI) cancer therapies such as GILOTRIF, GEFITINIB (Iressa) and ERLOTINIB (Tarceva), in patients with non-small cell lung cancer (NSCLC). Such mutations in the EGFR oncogene are present in the general population of patients with NSCLC at a frequency of approximately 7 to 23% in patients from the Western population of the USA and Europe, and 30 to 50% in Asian metastatic NSCLC patients. Excessive activation of EGFR has been found to be connected with cancer progression and poor prognosis. On the other hand, T790M mutation shows resistance to TKI therapy and therefore TKI therapy will not work on these patients. Thus, the detection of somatic mutations in the EGFR gene helps in choosing the appropriate targeted therapeutic regimens to increase the survival rate of cancer patients.
Detects 0.1% to 0.5% VAF mutant DNA out of wild-type DNA for targeted mutations
Powered by XNA technology, the QClamp® EGFR Mutation Detection Test has achieved a much higher analytical sensitivity than other commercial qPCR kits and other cancer gene mutation detection methods. QClamp® EGFR Mutation Detection Test can reliably detect 0.1% to 0.5% mutant DNA out of wild-type DNA for targeted mutations, providing a lower detection limit compared to similar assays available in the market due to robust enrichment of mutant sequences while suppressing amplification of wild-type sequences.
Supporting Data for QClamp® EGFR Mutation Detection Test
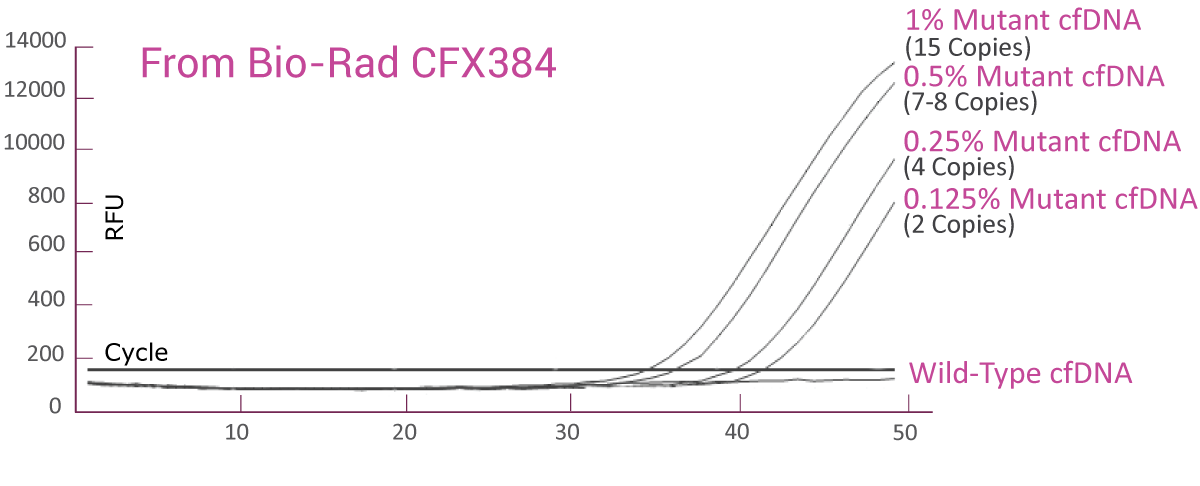
QClamp® EGFR Mutation Detection Test analytical sensitivity in plasma: detects as low as 0.125% VAF mutant DNA in a 5ng input
cfDNA extracted from plasma of a colorectal cancer patient with known EGFR T790M mutation was diluted in 5ng/μl of wild-type cfDNA. VAF of the sample was verified by Next-Generation Sequencing (NGS). The EGFR T790M assay was able to detect as low as 2 EGFR T790M mutant gene copies (0.125% VAF) in the background of wild-type cfDNA.
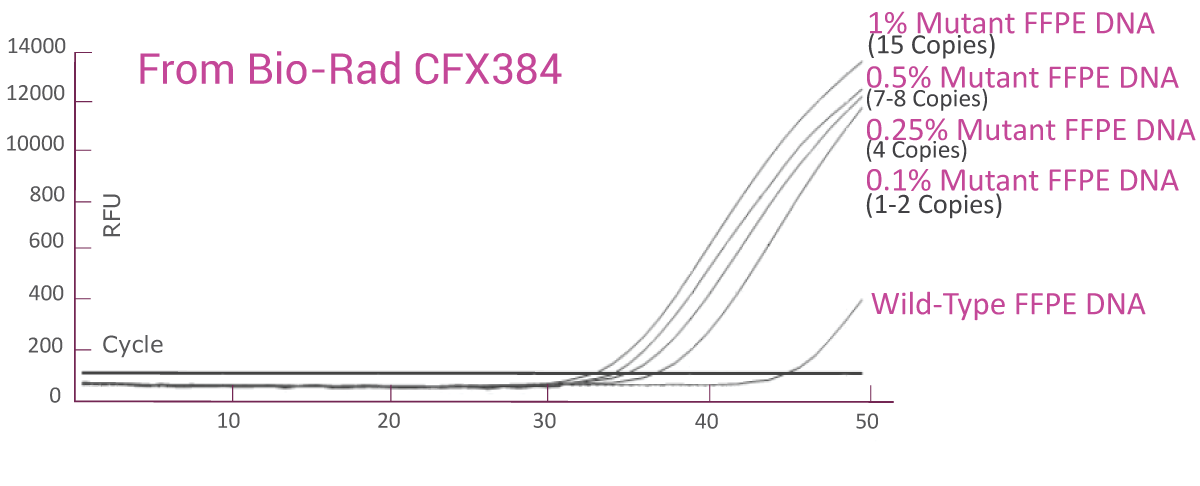
QClamp® EGFR Mutation Detection Test analytical sensitivity in FFPE: detects as low as 0.1% VAF mutant DNA in a 5ng input
DNA extracted from FFPE of a colorectal cancer patient with known EGFR T790M mutation was diluted in 5ng/μl of wild-type FFPE DNA. VAF of the sample was verified by NGS. The EGFR T790M assay was able to detect as low as 1.5 copies of EGFR T790M mutant (0.1% VAF) in the background of wild-type FFPE DNA.
Streamlined Workflow for QClamp® Gene Mutation Detection Tests

Step 1: DNA Isolation & Quantification
Extract DNA from FFPE or plasma using a commercial DNA extraction kit and measure the concentration using fluorometric analysis

Step 2: set up qpcr
Mix the assay reagents, load into PCR plate, add controls and extracted DNA ~ 30-60 minutes
Step 3: Amplification parameters
Enter amplification parameters on
qPCR instrument, load PCR plate
and start the run ~ 2.5 hours
Step 4: Data analysis
Determine the presence or absence
of mutations according to the Cq
value cutoffs ~ 15 minutes
Catalog Number
Pack size: 10-sample
CE Catalog #: DC-10-1038
Research-use-only (RUO) catalog #: DC-10-1038R
Pack size: 30-sample
CE Catalog #: DC-10-0012
Research-use-only (RUO) catalog #: DC-10-0012R
Detected Codons
Codons 719, 861, Ex19del, Ex20insASV and T790M, and L858R mutations
Sample Type
FFPE
Input DNA
5-10ng/Reaction
Validated Instruments
Roche LightCycler® 480, Bio-Rad CFX384 and ABI QuantStudio 5
Detection Channel
FAM; HEX
Detection Chemistry
TaqMan
Turnaround Time
Less than 4 hours
Stability
Stable for 12 months at -25 ℃ to -15 ℃
Most frequent EGFR mutations detected by QClamp® EGFR Mutation Detection Test
| Exon | Amino Acid Change | Nucleotide change | Cosmic No. |
|---|---|---|---|
| 18 | G719>A | c.2156G>C | 6239 |
| G719>S | c.2155G>A | 6252 | |
| G719>C | c.2155G>T | 6253 | |
| 19 | E746_A750del | c.2235_2249 del 15 | 6223 |
| E746_T751>I | c.2235_2252 > AAT | 13551 | |
| E746_T751del | c.2236_2253 del 18 | 12728 | |
| E746_T751>A | c.2237_2251 del 15 | 12678 | |
| E746_S752>A | c.2237_2254 del 18 | 12367 | |
| E746_S752>V | c.2237_2255>T | 12384 | |
| E746_A750del | c.2236_2250 del 15 | 6225 | |
| E746_S752>D | c.2238_2255 del 18 | 6220 | |
| L747_A750>P | c.2238_2248 >GC | 12422 | |
| L747_T751>Q | c.2238_2252 >GCA | 12419 | |
| L747_E749del | c.2239_2247 del 9 | 6218 | |
| L747_T751del | c.2239_2253 del 15 | 6254 | |
| L747_S752del | c.2239_2256 del 18 | 6255 | |
| L747_A750>P | c.2239_2248 TTAAGAGAAG>C | 12382 | |
| L747_P753>Q | c.2239_2258 >CA | 12387 | |
| L747_T751>S | c.2240_2251 del 12 | 6210 | |
| L747_P753>S | c.2240_2257 del 18 | 12370 | |
| L747_T751del | c.2240_2254 del 15 | 12369 | |
| L747_T751>P | c.2239_2251>C | 12383 | |
| K745_E749del | 2233_2247del15 | 26038 | |
| S752_I759del | 2253_2276del24 | 13556 | |
| E746_A750>IP | 2235_2248>AATTC | 13550 | |
| E746_T751>V | 2237_2252>T | 12386 | |
| E746_T751>IP | 2235_2251>AATTC | 13552 | |
| E746_S752>I | 2235_2255>AAT | 12385 | |
| E746_T751>VA | 2237_2253>TTGCT | 12416 | |
| E746_P753>VS | 2237_2257>TCT | 18427 | |
| L747_T751del | 2238_2252del15 | 23571 | |
| L747_S752>Q | 2239_2256>CAA | 12403 | |
| 20 | S768I | c.2303G>T | 6241/18486 |
| V769_D770insASV | c.2307_2308insGCCAGC... | 12376/26352 | |
| V769_D770insASV | c.2308_2309insCCAGCG... | 12426 | |
| T790>M | c.2369C>T | 6240 | |
| 21 | L858R2303 G>T | 2573_2574TG>GTS768I | 12429/6241 |
| L858>R | c.2573T>G | 6224 | |
| L861>Q | c.2582T>A | 6213 | |
Resources
Intended Use: QClamp® EGFR Mutation Detection Test is CE/IVD-certified. This product is available outside the USA for diagnostic use (CE/IVD) and research use (RUO). In the USA, this product is provided for research use only (RUO) and not for diagnostic use. DiaCarta ships the CE/IVD version to locations outside the USA and ships the RUO version within the USA. Please call 1-800-246-8878 or email order@diacarta.com if you have questions or specific needs.
Shipping Condition: QClamp® EGFR Mutation Detection Test will be shipped with dry ice. DiaCarta provides overnight delivery through FedEx Domestic Overnight Shipping Service for domestic shipments. For international shipment, DiaCarta provides 3-7 days in transit through FedEx International Priority Shipping Service. Please contact DiaCarta if you prefer to use your shipping carrier at order@diacarta.com.
